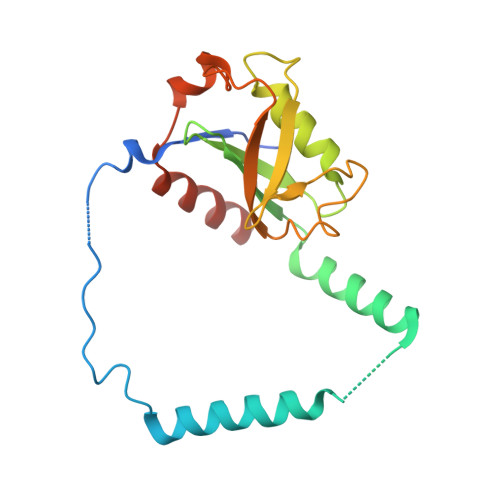
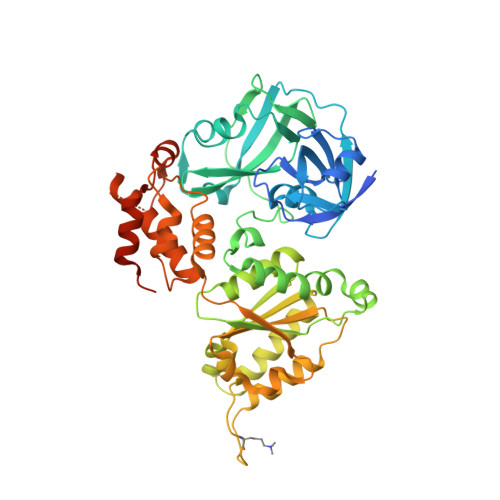
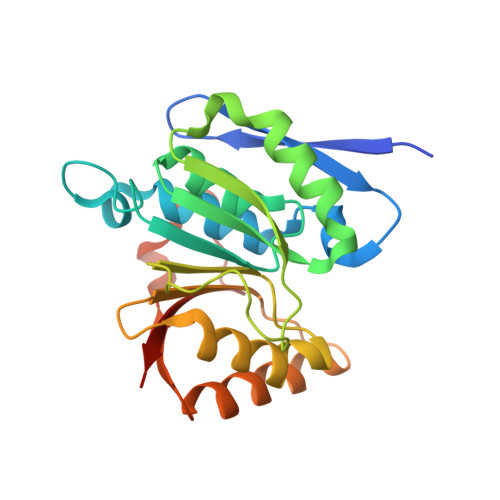
Structural remodeling of AAA+ ATPase p97 by adaptor protein ASPL facilitates posttranslational methylation by METTL21D.
Petrovic, S., Roske, Y., Rami, B., Phan, M.H.Q., Panakova, D., Heinemann, U.(2023) Proc Natl Acad Sci U S A 120: e2208941120-e2208941120
- PubMed: 36656859
- DOI: https://doi.org/10.1073/pnas.2208941120
- Primary Citation of Related Structures:
7OAS, 7OAT - PubMed Abstract:
p97 is an essential AAA+ ATPase that extracts and unfolds substrate proteins from membranes and protein complexes. Through its mode of action, p97 contributes to various cellular processes, such as membrane fusion, ER-associated protein degradation, DNA repair, and many others. Diverse p97 functions and protein interactions are regulated by a large number of adaptor proteins. Alveolar soft part sarcoma locus (ASPL) is a unique adaptor protein that regulates p97 by disassembling functional p97 hexamers to smaller entities. An alternative mechanism to regulate the activity and interactions of p97 is by posttranslational modifications (PTMs). Although more than 140 PTMs have been identified in p97, only a handful of those have been described in detail. Here we present structural and biochemical data to explain how the p97-remodeling adaptor protein ASPL enables the metastasis promoting methyltransferase METTL21D to bind and trimethylate p97 at a single lysine side chain, which is deeply buried inside functional p97 hexamers. The crystal structure of a heterotrimeric p97:ASPL:METTL21D complex in the presence of cofactors ATP and S-adenosyl homocysteine reveals how structural remodeling by ASPL exposes the crucial lysine residue of p97 to facilitate its trimethylation by METTL21D. The structure also uncovers a role of the second region of homology (SRH) present in the first ATPase domain of p97 in binding of a modifying enzyme to the AAA+ ATPase. Investigation of this interaction in the human, fish, and plant reveals fine details on the mechanism and significance of p97 trimethylation by METTL21D across different organisms.
Organizational Affiliation:
Max Delbrück Center for Molecular Medicine 13125, Berlin, Germany.